This is an old revision of the document!
Table of Contents
Compare different preparation protocols
Analysis description
ndeed, hEIDI has been developed to integrate the EDyP laboratory analysis pipeline :
Samples acquisition on mass spectrometer (Raw data)
Process of generated raw data into peak lists
Identification of proteins using Mascot search engine
Validation of the mascot identification result using IRMa
Export validated data into Mass Spectrometry Identification database
Exploit validated identification results by grouping identifications at the whole database level to remove redundancy or comparing multiple experiments …
In this tutorial, we will work with two biological replicates (named “Replicate1” and “Replicate2”).
Each replicate is aliquoted in three parts and we realize a different preparation protocol for each part (“Ultrafiltration”, “Proteominer” and “TCA Precipitation”).
Each aliquote is then deposed onto a SDS-Page gel.
When separating is done, bands of interest are extracted from the gel and analysed using the EDyP laboratory analysis pipeline:
HPLC-MSMS → Mascot(protein identification) → IRMa(validation) → MSIdb(database with valid identification results)
Each gel band will result into one identification result (named “F093496.dat” for example). Then, the analysis of a whole aliquote will generate many identification results.
...Project ABC
...Replicate1
.Ultrafiltration
.F093496.dat
.F093497.dat
...
.Proteominer
...
.TCA Precipitation
...
...Replicate2
.Ultrafiltration
...
.Proteominer
...
.TCA Precipitation
...
Project creation in hEIDI
Please follow the steps in the getting start to setup the MSI db connection (Mass Spectrometer Identification database), create an hEIDI project and open a working session on the MSI db.
Check the filter parameters used in identification results
 verifier les paramètres de filtres des identification (how to view ident properties + .. view filter )
verifier les paramètres de filtres des identification (how to view ident properties + .. view filter ) 
How to check filter parameters used in idenitification results
Structure your experiment as a context tree
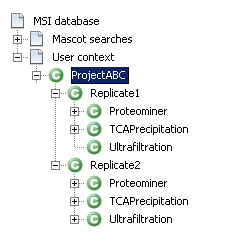 Now you need to build a context hierarchy and dispatch your identification results to map the project structure described in the first section.
Now you need to build a context hierarchy and dispatch your identification results to map the project structure described in the first section.
See the following links to know what is a context and how to create a context hierarchy.
Execute protein grouping for every contexts
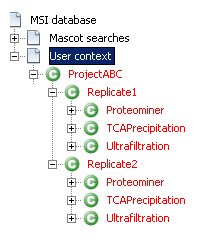 Protein grouping is the next step. Execute the protein grouping algorithm for all the User contexts in a specific order.
Protein grouping is the next step. Execute the protein grouping algorithm for all the User contexts in a specific order.
- Start with the contexts that directly group the identification results: “Ultrafiltration”, “Proteominer” and “TCA Precipitation”. Do that for the two replicates.
- Then continue with the contexts “Replicate1” and “Replicate2”.
- And finally, end with the root context named “ProjectABC”
If everything has worked correctly, all the context names are now in red.
Browse the context tree and export it
Check reproducibility of results
You may want to compare results from the same preparation protocol in the two replicates.
Compare two contexts with each other and export comparison results
 a deplacer
a deplacer 
How to compare contexts each other.
See the following link to understand what is context comparison.
Find the optimal preparation protocol
You may want to compare protocols each other for a given replicate to find the “best” one.
Compare a parent context with their child contexts and export comparison results
 a deplacer
a deplacer 
- Show covering diagram
- Export results
See the following link to understand what is context comparison.

