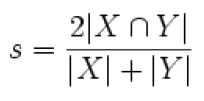This is an old revision of the document!
Table of Contents
Context comparisons
Definition
The purpose is to search for similar protein groups, i.e. protein groups:
- having the same set of proteins
- identified by a same set of peptides
The Dice coefficient is used to measure similarity (result ranges from 0 to 1).
Compare several contexts to their union
The purpose is to select contexts to compare, then to compare them to their union, i.e. a parent context, which contains selected children contexts, and for which peptide/protein grouping has been executed.
Is there a need to create a "virtual" union parent context ?
There are 2 possibilities, either the parent union already exists or the parent union needs to be created.
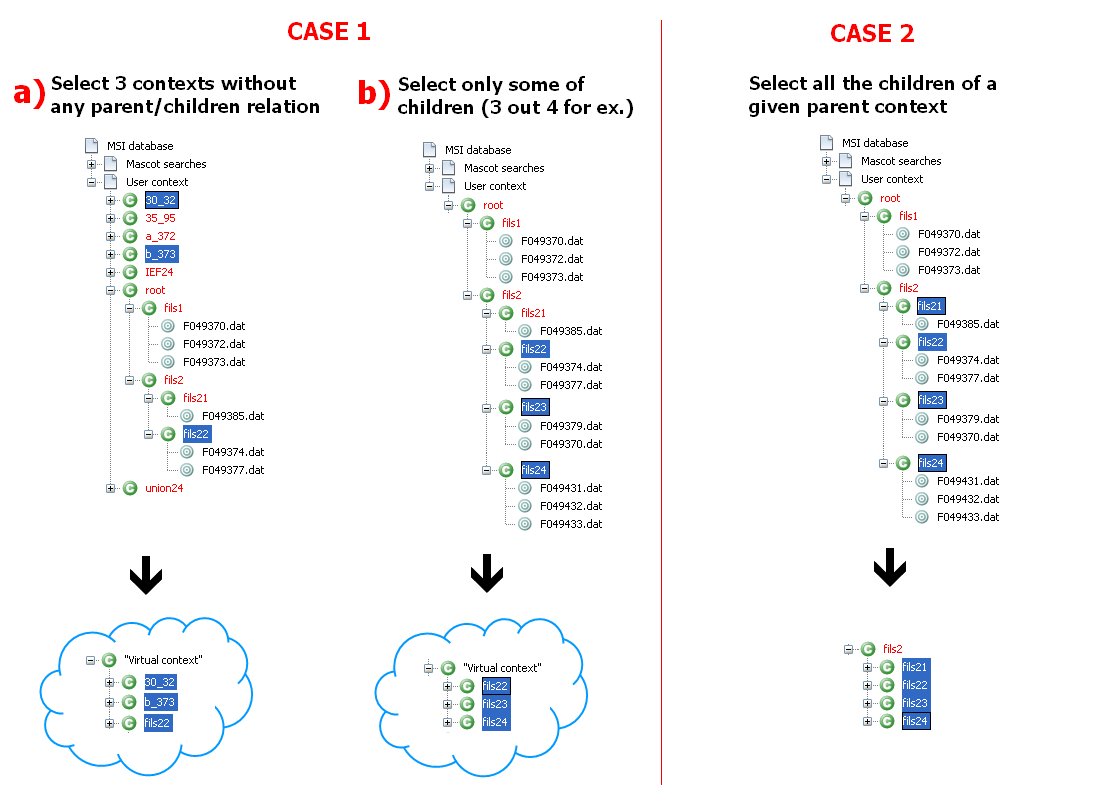
In the image below:
- the union parent must be created in case 1: it is called virtual as it is only created in-memory for the process
- the union parent already exists in case 2
Compare union parent context to each children
The algorithm will then compare every protein groups of the union parent context to every protein groups of each children context in order to find the best alignment.
Comparing two protein groups each other means:
- Checking the similarity of their peptide sets
- Checking the similarity of their protein sets (sameset and subset)
Case 1: if a Protein Group (context1) is compared to the 2 following Protein Groups (Context2), PG2 will be preferred
| PG1 | PG2 | |
|---|---|---|
| peptideSimilarity | 0.2 | 0.5 |
| proteinSimilarity | 0.4 | 0.6 |
| AllProtSimilarity | - | - |
Case 2: if a Protein Group (context1) is compared to the 2 following Protein Groups (Context2), PG2 will be preferred
| PG1 | PG2 | |
|---|---|---|
| peptideSimilarity | 0.5 | 0.5 |
| proteinSimilarity | 0.4 | 0.6 |
| AllProtSimilarity | - | - |
Case 3: if a Protein Group (context1) is compared to the 2 following Protein Groups (Context2), PG2 will be preferred
| PG1 | PG2 | |
|---|---|---|
| peptideSimilarity | 0.5 | 0.5 |
| proteinSimilarity | 0.6 | 0.6 |
| AllProtSimilarity | 0.5 | 0.6 |
Case 4: if a Protein Group (context1) is compared to the 2 following Protein Groups (Context2), PG2 will be preferred
| PG1 | PG2 | |
|---|---|---|
| peptideSimilarity | 0.0 | 0.0 |
| proteinSimilarity | 0.6 | 0.8 |
| AllProtSimilarity | - | - |
Details about comparison results
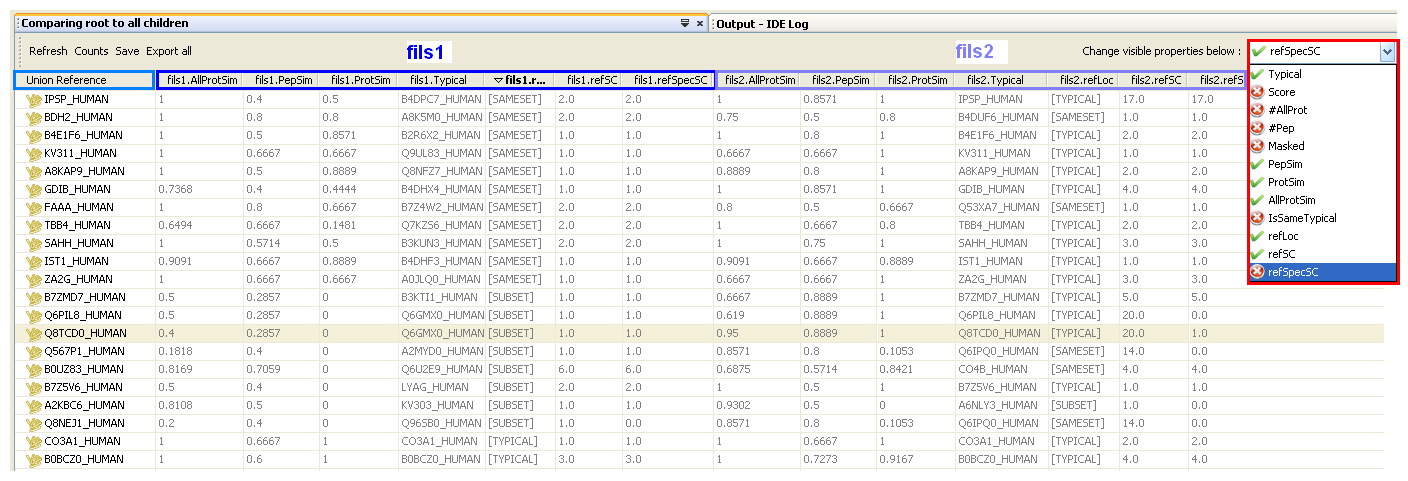 When the comparison algorithm execution is finished, a window displays on the screen. For each compared context a default set of properties are displayed. You can change them by selecting/deselecting items in the droplist (right side).
When the comparison algorithm execution is finished, a window displays on the screen. For each compared context a default set of properties are displayed. You can change them by selecting/deselecting items in the droplist (right side).
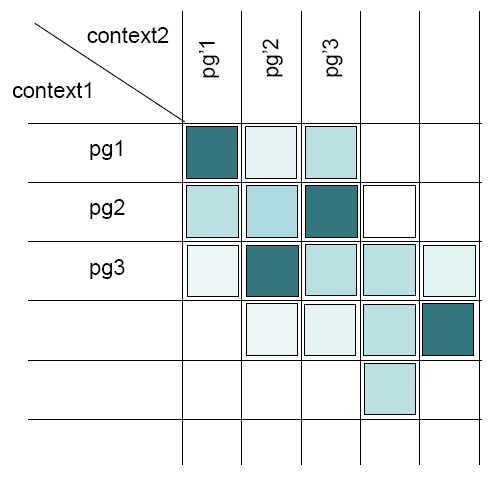
In the short view, the “Typical reference” column lists the protein groups of the parent context. There are as much columns as there are direct child contexts for this parent context. These columns display the value of the parameter selected in the droplist (“#peptides”, “Typical”, “Score”, “Pep. similarity”, “Prot. similarity”, “All Prot. similarity”).
 Different criteria to compare protein groups:
Different criteria to compare protein groups:
- “ProtSim”: similarity on the samesets of proteins of the two protein groups
- “AllProtSim”: similarity on the samesets and subsets of proteins of the two protein groups
- “PepSim”: similarity on peptides of the two protein groups
- “isSameTypical”: indicates if the typical protein is the same between the two protein groups
- “Typical”: the accession of the master protein group
- “Score”: score for the protein group
- “#Pep”: number of peptides for the protein group
- “#AllProt”: number of proteins in the protein group. 'X(Y,Z)' means a total of X proteins for the whole protein group (X=Y+Z) with Y sameset proteins and Z subset proteins
If Spectral Count algorithm has been run previously, Spectral Count properties will also be available.
When comparing two protein groups using “Protein” and “Peptide” similarity criteria, you can get four main situations:
| Low Protein similarity | High Protein similarity | |
|---|---|---|
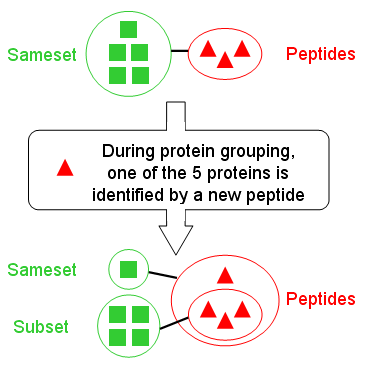
| Low Peptide similarity | Proteins groups are not alike | We have identified the same proteins but with different peptides (rare) |
| High Peptide similarity | See image (on the left) to have an example of how to get this situation. It's usefull to check the “All Protein” similarity to have an information on proteins in subset | protein group are alike |
Compare two contexts with each other
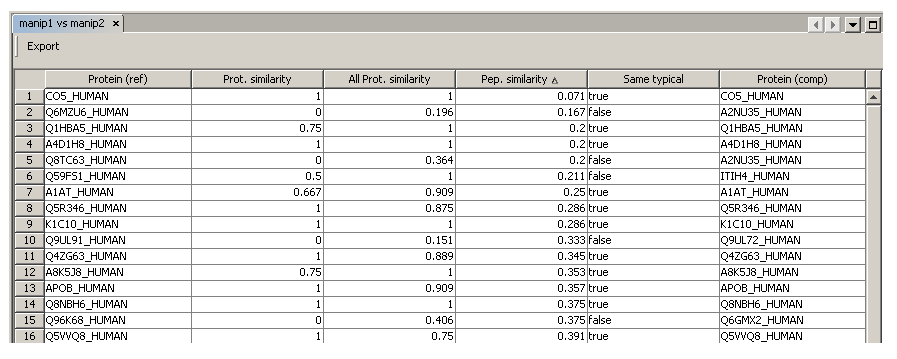
The most left column (“Protein (ref)”) displays protein groups from the first context. The most right column (“Protein (comp)”) displays protein groups from the second context.
For each protein group of the first context, the table proposes the most “similar” protein group in the second context (“Protein similarity”, “All protein similarity”, “Peptide similarity”,“Same typical”).