userguide:retentiontimealignment
This is an old revision of the document!
Retention Time Alignment
Here are the major steps followed by the Retention Time algorithm:
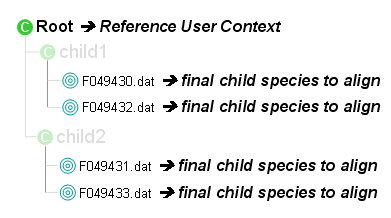
- Collect species from the reference User context and predict their retention time using an external utility
- For each identification gathered under the reference User context do:
- Collect species to align from the identification
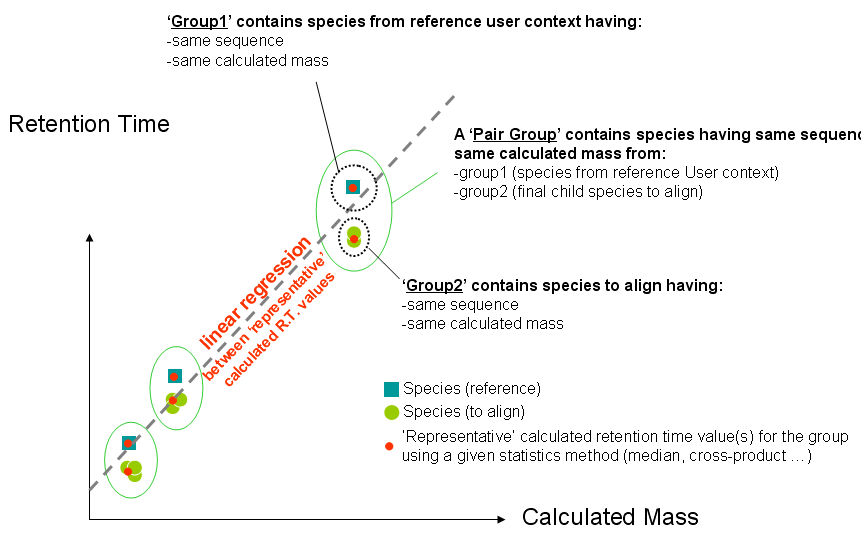
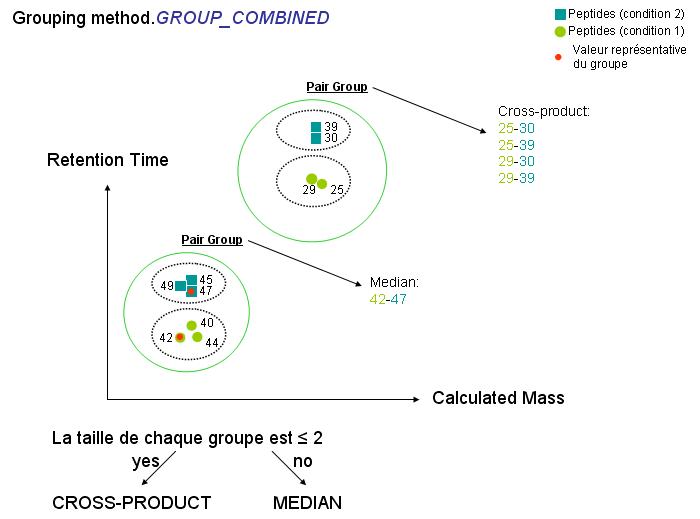
- Constitute several pair groups between collected reference species & species to align. A pair group contains 2 groups of species having same sequence & calculated mass (group1 has species from reference User context and group2 has species to align from identifications).
- Compute one (or several) representative value(s) for group1 & group2 for each pair group
- Compute linear regression between representative values
- Store linear regression coefficients
In more details…
- Species Retention Time of the reference UserContext are predicted with NETPrediction v2.2.3378 utility using Kangas method (click here for more details). NETPrediction utility only uses species sequences to predict a Normalized Elution Time (NET) value. First, a list of 'reference' species is built with the below criteria, then the corresponding sequences are exported in order to be used by the NETPrediction utility:
- The reference species list doesn't contain any species with PTMs
- If several species exist with redundant sequences, the best score species is retained
- All the final child species, i.e. species in identifications, are collected.
userguide/retentiontimealignment.1280740658.txt.gz · Last modified: 2010/08/02 11:17 by 132.168.74.230