This is an old revision of the document!
Table of Contents
Protein grouping
Algorithm
Step 1 - Peptide grouping
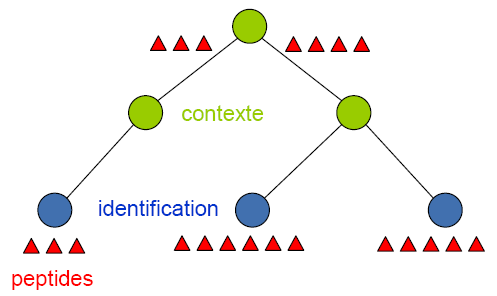
Peptides from different identifications - attached directly or indirectly to a context - are grouped.
Peptides must have same sequence and same experimental mass to be grouped.
Peptide grouping results in new peptides attached to the parent context and having child peptides
Step 2 - Protein grouping
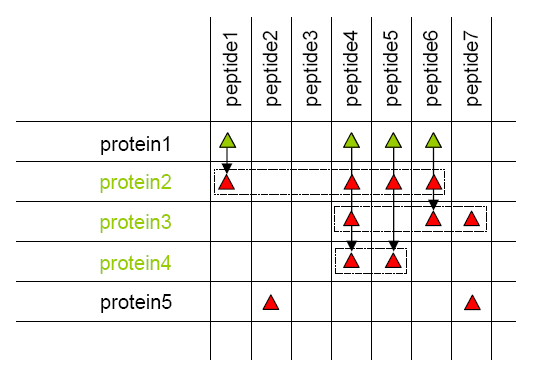
Peptides from a same protein or a same protein set are grouped.
Protein grouping results in new groups of proteins and peptides, attached to the parent context.
![]() You need to be carefull when grouping proteins within a tree of contexts. Let's take the following example:
You need to be carefull when grouping proteins within a tree of contexts. Let's take the following example:
Rootnode
|_ Context1
|_ F085255.dat
|_ F085256.dat
|_ F085257.dat
|_ Context2
|_ F085258.dat
|_ F085259.dat
It's possible:
- case 1 - to group proteins at the
Rootnodelevel, hEIDI will then group proteins from all the identification results, or - case 2 - to group proteins starting from the leaf contexts (
Context1andContext2), then ending with theRootnode.
At present, when launching the protein group algorithm, you can tell hEIDI to filter some proteins and/or peptides. For example, if you decide to filter proteins with a number of peptides lower than 2, it is important to understand that doing this may give different results in cases 1 & 2.
Rootnode
|_ Context1
ProtA (pep1, pep2)
|_ Context2
ProtA (pep1, pep5)
In the very trivial example above, you filter, when grouping proteins in each leaf context, the proteins with a number of peptides lower than 2. This will remove ProtA and then, when grouping proteins in the Rootnode context, ProtA will not appear in the final result.
But in case 1, when grouping/filtering proteins at the Rootnode level, ProtA will gain one peptide more (ProtA will be identified by 3 peptides instead of 2) and so will appear in the final result.